Accessing Pluto data programmatically
Last updated: October 20, 2025
Use the Pluto API to fetch data from your projects and experiments, and integrate Pluto with your own scripts and apps
Written by Daniel Shin
To perform your own custom analysis on data stored in Pluto, or to integrate Pluto data into your own apps or scripts, you can use the Pluto API token to fetch data programmatically.
To fetch data, you'll typically need 3 pieces of information:
Pluto API token
Pluto experiment ID (e.g. PLX000327)
Plot ID (if fetching results produced from a specific analysis)
Creating your API token
If you are accessing data programmatically for the first time, you'll need to generate a new API token. Navigate to your Account page and click the "+ API token" button.
A new API token will be generated, and can then be copied or viewed.
This API token can be used to fetch data programmatically using a curl request or Pluto's R package.
R package
1. Set the PLUTO_API_TOKEN environment variable
Copy the API token that you created on your Account page and store it in an environment variable called PLUTO_API_TOKEN. To do this, create a .Renviron file containing:
PLUTO_API_TOKEN="<YOUR API TOKEN>"
2. Source the latest version of the R package
# Load required dependencies
library(httr)
library(jsonlite)
library(rjson)
# Print PLUTO_API_TOKEN to confirm that it's in your environment
#Sys.getenv('PLUTO_API_TOKEN')
# If PLUTO_API_TOKEN is blank, make sure the API key is in .Renviron
# and restart your R session, then rerun Sys.getenv('PLUTO_API_TOKEN')
# Import the pluto_read() function
devtools::source_url("https://github.com/pluto-biosciences/pluto-sdk-R/blob/main/pluto.R?raw=TRUE")
3. Fetch sample data for an experiment
sample_data <- pluto_get_experiment_data(experiment_id ='PLX140206'
, table_type = 'sample')
# All 12 rows of the sample data were fetched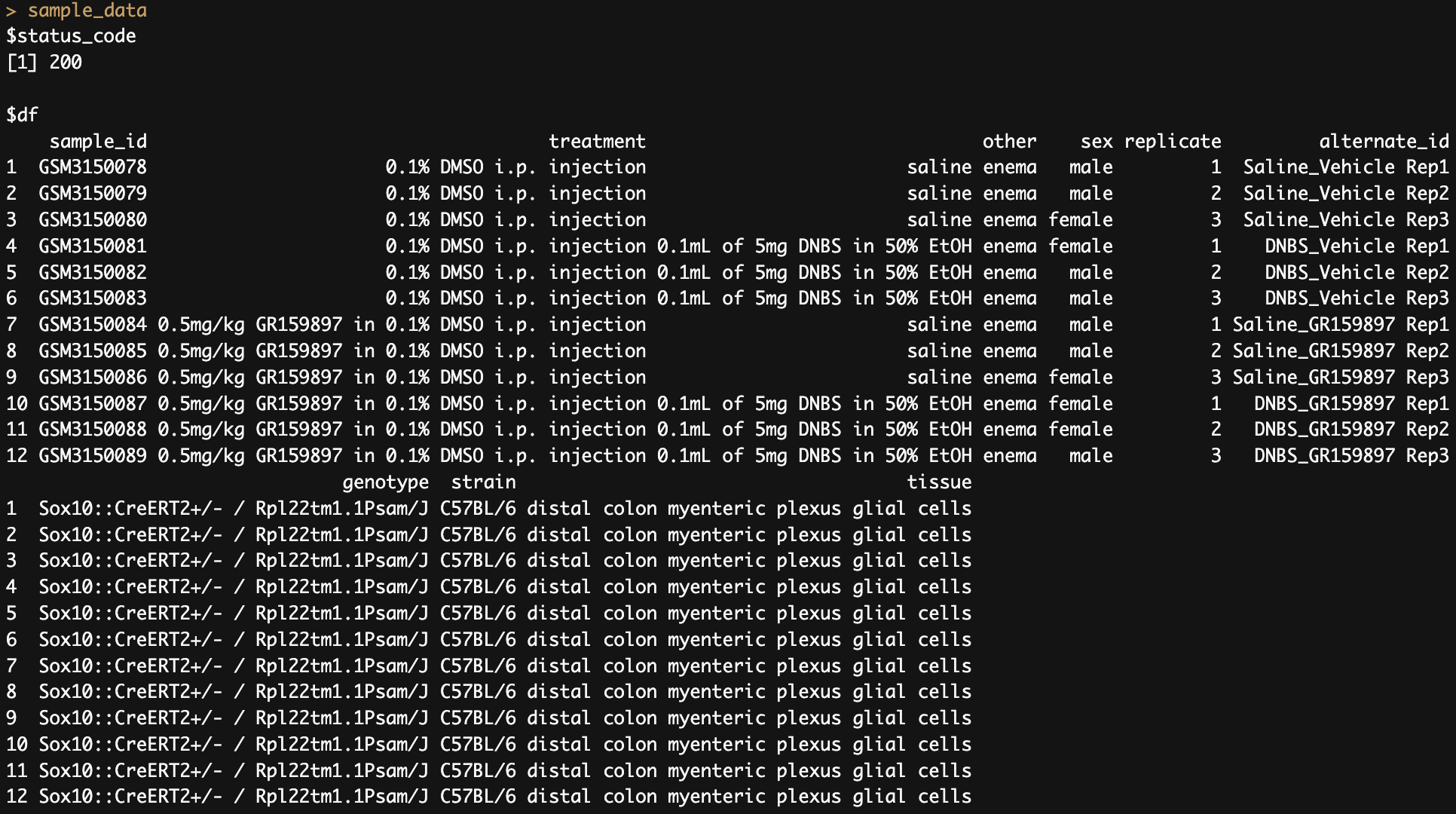
4. Fetch assay data for an experiment
assay_data <- pluto_get_experiment_data(experiment_id ='PLX140206'
, table_type = 'assay')
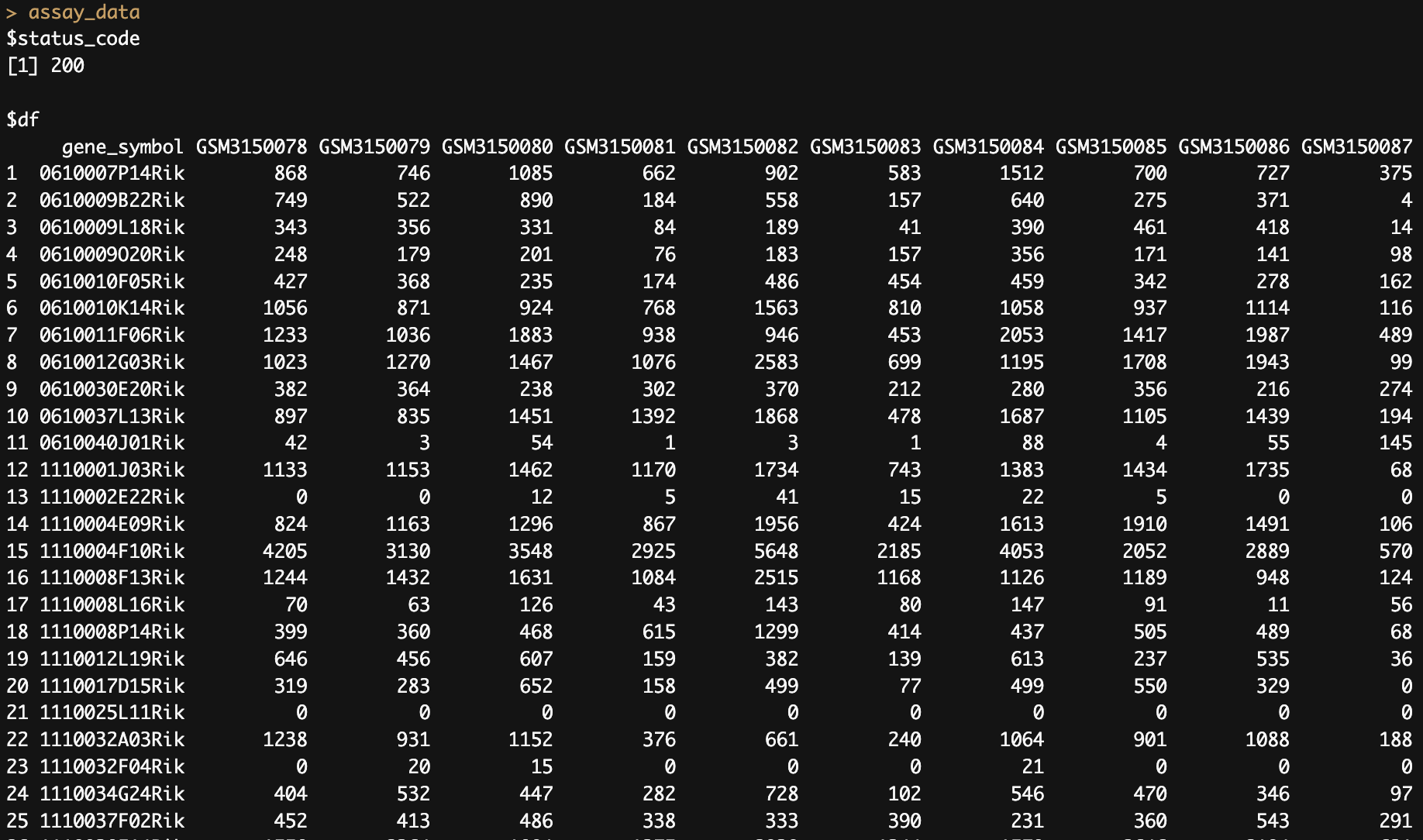
5. Fetch the results table for a specific experiment & analysis
Obtain the plot ID from the Methods Modal on any plot:
Use the plot ID in the pluto_get_results() function to fetch the underlying results:
deg_table <- pluto_get_results(experiment_id = "PLX140206",
plot_id = "386b3bbf-fc86-4071-88c2-6340634cdc62")
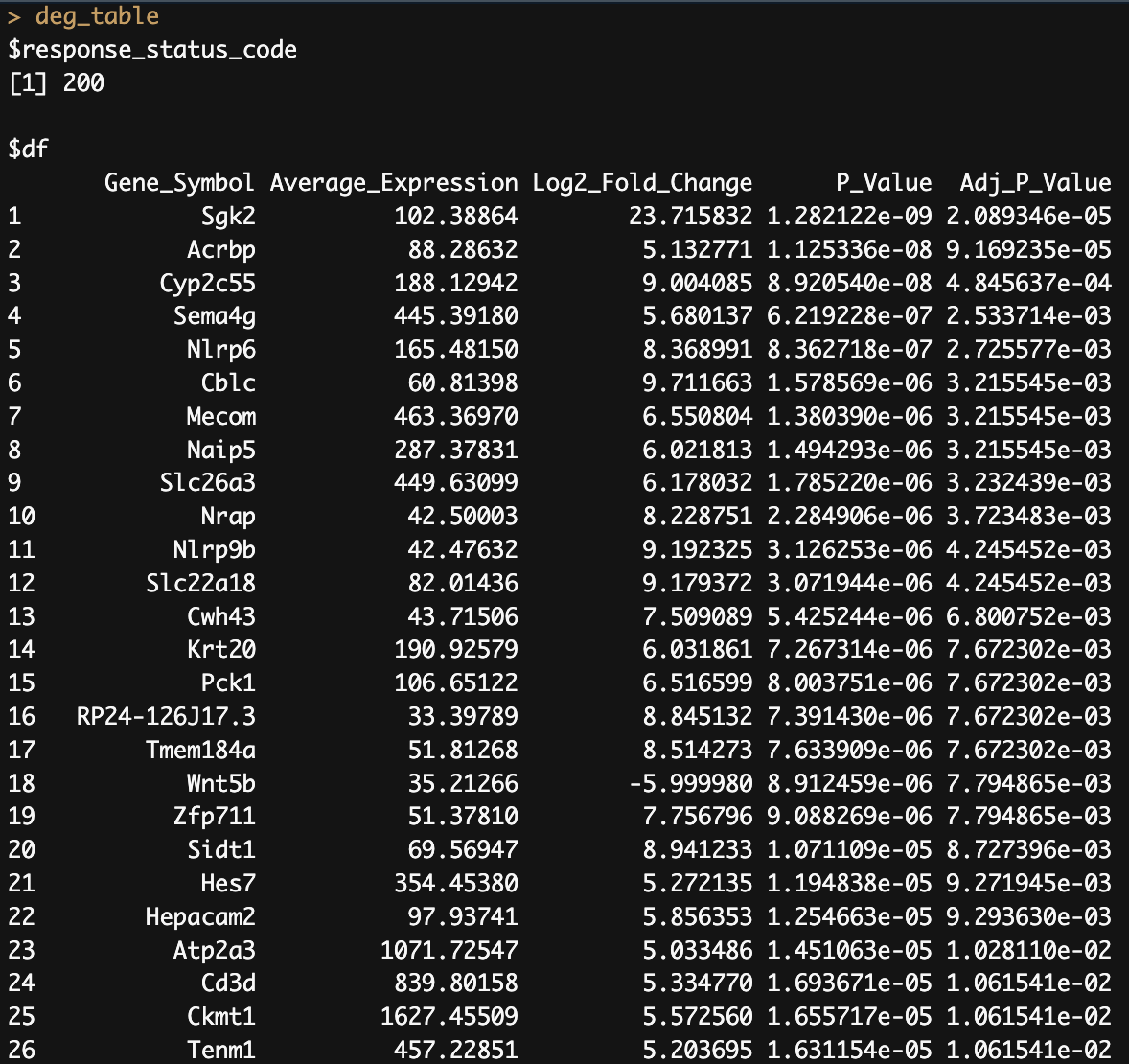
Python example
import requests
import pandas as pd
from scipy.stats import zscore
# API token for authenticating request
headers = {"Authorization": "Token <YOUR_API_TOKEN>"}
# Request parameters
proj_id = "<PROJECT_ID>"
expt_id = "<EXPERIMENT_ID>"
plot_id = "<PLOT_ID>"
limit = "10000"
# Endpoints
project_experiments_endpoint = "https://api.pluto.bio/lab/projects/" + proj_id + "/?limit=" + limit
assay_data_endpoint = "https://api.pluto.bio/lab/experiments/" + expt_id + "/assay-data/?limit=" + limit
sample_data_endpoint = "https://api.pluto.bio/lab/experiments/" + expt_id + "/sample-data/?limit=" + limit
all_plots_endpoint = "https://api.pluto.bio/lab/experiments/" + expt_id + "/plots/?limit=" + limit
plot_data_endpoint = "https://api.pluto.bio/lab/experiments/" + expt_id + "/plots/" + plot_id + "/data/?limit=" + limit
# Fetch assay data for an experiment
assay_data_df = pd.DataFrame(requests.get(assay_data_endpoint, headers=headers).json()["items"])
# Fetch sample data for an experiment
sample_data_df = pd.DataFrame(requests.get(assay_data_endpoint, headers=headers).json()["items"])
# Fetch plot results for a single analysis
results_df = pd.DataFrame(requests.get(results_endpoint, headers=headers).json()["items"])
# Fetch all plots for an experiment
plots_response = requests.get(all_plots_endpoint, headers=headers)
plots_df = pd.DataFrame(plots_response.json()).sort_values("analysis_type")
CURL requests
CURL requests must provide your PLUTO_API_TOKEN and experiment_id and data_type ("sample", "assay", or "results").
Deleting your API token
You can delete your API token by clicking the "Delete token" button. Caution: deleting your API token will immediately remove any access associated with that token. This means that any apps or scripts that were using this token will no longer be able to fetch data.