Entity databases
Last updated: September 3, 2025
Build flexible databases to capture, manage, and standardize metadata about in vitro and in vivo models, patient cohorts, tissue ontologies, compounds, and any other entities your organization needs to reference.
Overview
R&D and translational teams frequently manage growing lists and tables of "things," from compounds, to mouse models, to patient metadata, that the team needs to reference.
Traditionally, these lists are maintained in spreadsheets or buried across different documents, which can lead to version control issues, mismatched identifiers, and duplicated work.
Entity databases in Pluto solve this problem by:
Providing a centralized, structured place to store reference metadata
Ensuring consistency across experiments (e.g., the same patient ID or compound identifier is used everywhere)
Making it easy to cross-reference entities across modalities (e.g., RNA-seq and ChIP-seq that use the same compound)
Creating a new entity database
On the Entity databases page, select the blue “+ New database” button and give your database an informative name and description. Select or define a type for the database (e.g. Cell line, Patient, Mouse, Compound, Organ-chip, etc.)
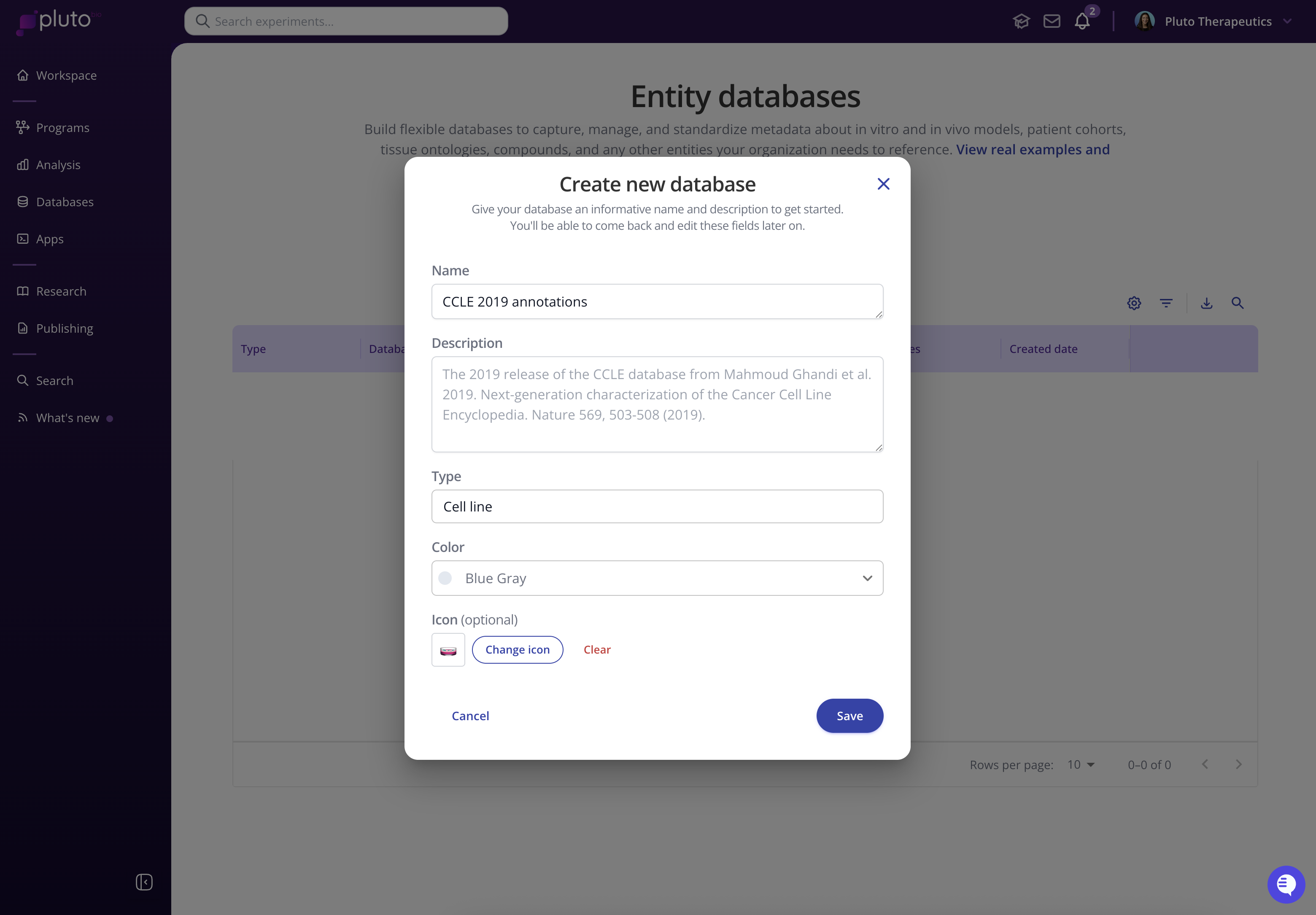
Defining the database schema
When you click into your newly created database, there are two methods for defining its schema, or the set of fields/columns in your database:
Define the schema by creating fields individually
Import a CSV file and use its column headers to define the database schema

Adding a field to the schema individually
Select "Define schema" and choose "Add field". Give your field a name (e.g. "ModelID") and select the type of values that it will contain.
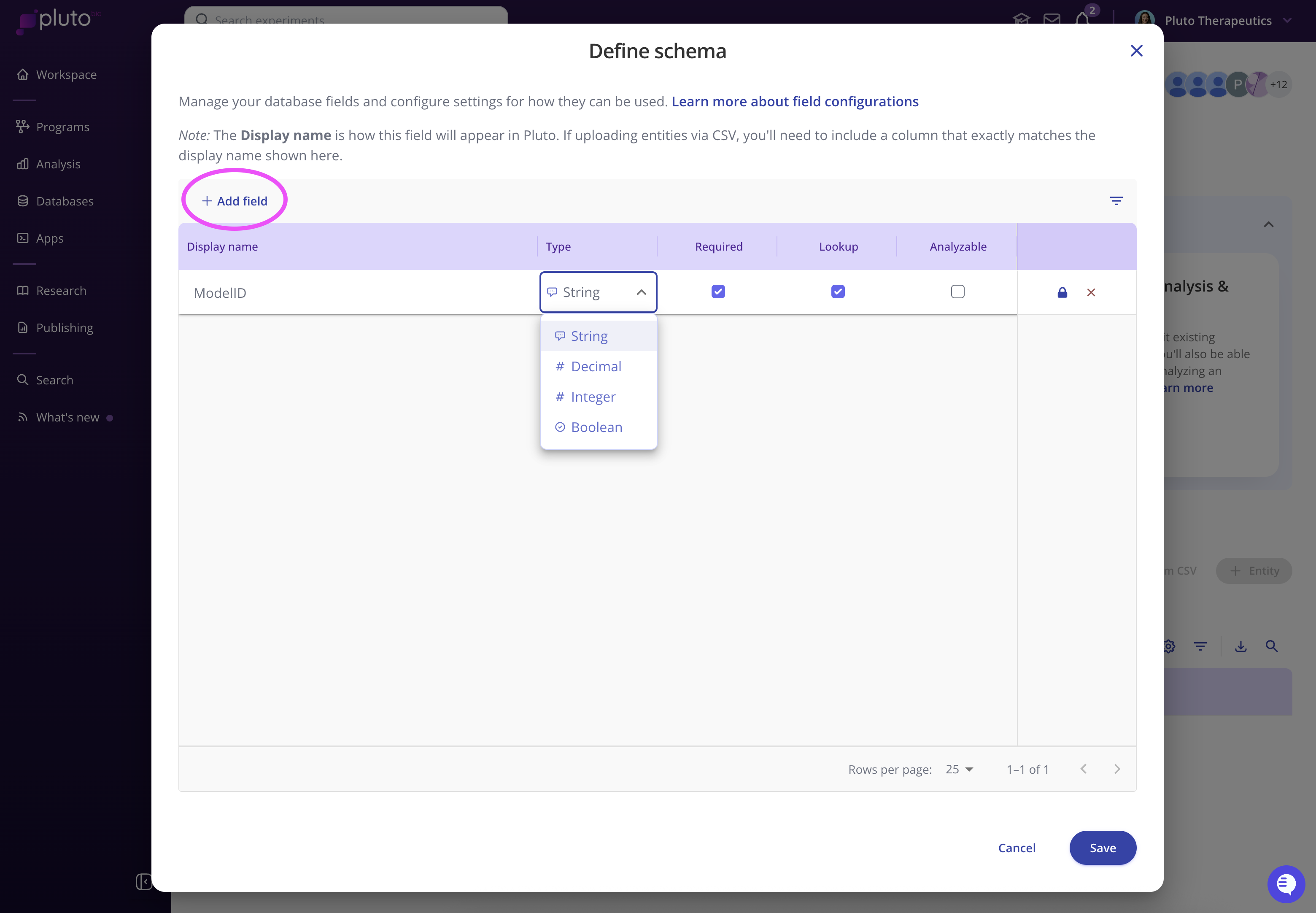
Use the checkboxes to configure how you want a field to be used:
Required- This field will be required to be non-empty for every entity record in the database.Lookup- Use this checkbox if you intend to use this field as an identifier for mapping to experimental data sets.Analyzable- These fields will be available on experimental data sets when performing analysis. Typically, this will include fields that you would group samples by for visualization, such as tissue type or response.
Continue adding any fields (columns) that you intend to capture for each entity. You can also make edits to these fields later on.
Example for a cell line database: Name, Source, Doubling time, Tissue of origin, TP53 Mutation Status
Example for a patient cohort database: Patient ID, Age, Sex, Diagnosis, Treatment arm, Outcome
Adding & editing database entities
Populate your database with entries (records). Each entry is an entity instance, like “MCF-7” (cell line), “DrugX” (compound), or “Patient_001.”
Once your database is created, you can link entities directly in your sample annotation tables when running analyses in Pluto.
Examples & Inspiration
Patient cohorts
TCGA BRCA patients – clinical metadata
Institutional patient cohorts with treatment response data
Cell lines
CCLE – 2019 published version
Your lab’s internal immortalized lines with custom fields
Animal models
Jackson Laboratory mouse models
Crown Bio PDX models
Compounds
Approved drugs in your therapeutic area
Proprietary small-molecule libraries
Tissue / Ontologies
Standardized tissue definitions for consistent annotation
Human organoids and organ-chip models
Other powerful entities
Antibodies and reagents
Protocols, assay platforms, sequencing prep, instrument
Clinical visit timepoints and dosing regimens
Best practices for entity databases
Start broad, refine later - don't worry about defining every field in the schema up front. Begin with the key fields you need, then expand as your team identifies additional metadata.
Leverage across assays. Create databases you know will be referenced repeatedly (patients, compounds, cell lines).
Think globally. If multiple groups in your org will use the same definitions, capture them once in a central entity database.
⚡ Tip: Many teams find it useful to start with a Cell line database (like CCLE) or a Patient cohort database (like TCGA) as inspiration. You can import public datasets or build your own from scratch.